Design of QTY Retinylidene Proteins
In 2025, I conducted bioinformatics research on designing soluble versions of retinylidene proteins, a type of protein essential for human vision, and published a paper in QRB Discovery by Cambridge University Press.
While studying the QTY code, a protein design method devised by Prof. Shuguang Zhang and his team, I began thinking about designing soluble rhodopsins. Having studied ophthalmology and neuroscience since eighth grade, I had accumulated much knowledge about the visual system. I knew that rhodopsin, which was an insoluble protein important for vision, is implicated in certain cases of retinitis pigmentosa, the disease that my grandmother has. Soluble versions of rhodopsin could make it easier to explore the relationship between its structure and function, potentially accelerating the development of treatments. I then went on to explore other proteins and ultimately selected 9 human opsins and 3 microbial opsins to study. Rhodopsin remained the primary focus, as I examined all the proteins structurally but investigated rhodopsin functionally.
I conducted research from January to July.
In January and February, I learned molecular biology and did background research, reading over 150 papers.
In late February, I designed the proteins with the QTY code and investigated their characteristics and structures.
In March and April, I ran molecular dynamics simulations and verified the results.
In May, I made the figures and wrote the paper.
In June and July, I managed the publication process, which included revising the paper and responding to peer reviews.
Through this research project, I learned about many things:
Principles of molecular biology, especially the aspects related to protein structures and functions;
The mathematics and physics of molecular dynamics simulations;
Tools for designing water-soluble QTY analogues of proteins: online QTY server and Protter;
Tools for protein structure prediction: AlphaFold3;
Tools for examining protein structures and surface properties: PyMOL, ChimeraX, and Avogadro;
Tools for molecular dynamics simulation: GROMACS, CHARMM-GUI, CGenFF, and others;
Tools for visualization: Python, especially Matplotlib.
Scientific conventions in academic writing;
The process of publication.
Abstract
Retinylidene proteins are retinal-binding light-sensitive proteins found in organisms ranging from microbes to human. Microbial opsins have been utilized in optogenetics, while animal opsins are essential for vision and light-dependent metabolic functions. However, retinylidene proteins have hydrophobic transmembrane (TM) domains, which makes them challenging to study. In this structural and functional bioinformatics study, I use the QTY (glutamine, threonine, tyrosine) code to design water-soluble QTY analogues of retinylidene proteins, including nine human and three microbial opsins. I provide superpositions of the AlphaFold3-predicted hydrophobic native proteins and their water-soluble QTY analogues, and experimentally determined structures when available. I also provide a comparison of surface hydrophobicity of the variants. Despite significant changes to the protein sequence (35.53–50.24% in the TM domain), protein characteristics and structures are well preserved. Furthermore, I run molecular dynamics (MD) simulations of native and QTY-designed OPN2 (rhodopsin) and analyze their response to the isomerization of 11-cis-retinal to all-trans-retinal. The results show that the QTY analogue has similar functional behavior to the native protein. The findings of this study indicate that the QTY code can be used as a robust tool to design water-soluble retinylidene proteins. These have potential applications in protein studies, therapeutic treatments, and bioengineering.
Publication Information
Pan, S. (2025). A structural and functional bioinformatics study of QTY-designed retinylidene proteins. QRB Discovery, 6, e20. https://doi.org/10.1017/qrd.2025.10009
The QTY code is a method for designing soluble versions of membrane proteins. To use an analogy, it is like a magic spell that lifts trees from the ground, letting them float in midair with their roots intact. Normally, membrane proteins ("trees") are stuck in the cell membrane ("ground") because part of them ("roots") fears water ("air"). The QTY code changes this water-fearing part ("root"), freeing the protein and making it water-soluble ("afloat in the air"). Here is a more technical explanation:
The transmembrane segments of membrane proteins are hydrophobic in nature. This leads to a challenge to study their structure and function. Traditionally, detergent must be used to purify them. Zhang et al. considered a different approach, that is, via systematic soluble protein design. There exist structural similarities, as can be observed on high-resolution electron density maps, between certain hydrophobic and hydrophilic amino acids: leucine (L) and glutamine (Q), isoleucine (I) or valine (V) and threonine (T), and phenylalanine (F) with tyrosine (Y). This fact enables systematic replacement of L with Q, I/V with T, and F with Y in all transmembrane segments of the membrane protein, which Zhang et al. named the QTY code. QTY analogues are less hydrophobic than native membrane proteins. Although their amino acid sequences are significantly changed, they still exhibit relatively preserved structure, isoelectric points (pI), and molecular weights (MW) compared to the native membrane proteins.
I selected a variety of retinylidene proteins (opsins), with 9 animal (human) ones and 3 microbial ones. Some of them are better studied and implicated in diseases, while others are less well studied. In either case, QTY-designed soluble versions show promise in facilitating futures studies and applications. This is a brief introduction to all these proteins:
Animal opsins are a type of class A GPCR (G-protein-coupled receptor), which are characterized by seven transmembrane domains, the NPxxY motif, and activation of G proteins through the outward movement of TM6. Almost all animal opsins include a lysine residue that forms a Schiff base link with the retinal. When retinal absorbs a photon, it isomerizes, usually changing from 11-cis to all-trans. The subsequent conformational changes of the protein are well studied using bovine rhodopsin, which changes from dark state to BATHO state, then LUMI, META I, and META II. Proton transfer plays an important role in this process. Afterward, the protein bleaches and returns to the dark state, completing what is called the photocycle. As the protein conformation changes to META, TM6 moves characteristically outward, activating the G protein. In photoreceptor cells, the G protein, transducin, activates a phosphodiesterase, which hydrolyzes cGMP into GMP, decreasing the activity of cGMP-gated cation channels and hyperpolarizing the cell.
In this study, I select nine opsins expressed in the human nervous system: OPN1MW (UniProt ID: P04001), OPN1LW (UniProt ID: P04001), OPN1SW (UniProt ID: P03999), OPN2 (UniProt ID: P04001), OPN3 (UniProt ID: Q9H1Y3), OPN4 (UniProt ID: Q9UHM6), OPN5 (UniProt ID: Q6U736), RGR (UniProt ID: P47804), and RRH (UniProt ID: O14718). They belong to several evolutionarily distinct families of animal opsins.
OPN1MW (medium-wave-sensitive opsin 1), OPN1LW (long-wave-sensitive opsin 1), and OPN1SW (short-wave-sensitive opsin 1) are expressed in retinal cone photoreceptors and are responsible for color vision. Certain variants of OPN1MW, OPN1LW, and OPN1SW, respectively, cause deuteranopia, protanopia, and tritanopia, which are different types of color blindness. The absence of both functional OPN1MW and OPN1LW causes blue cone monochromacy, an X-linked congenital cone dysfunction syndrome, and cone dystrophy 5, an X-linked cone dystrophy.
OPN2 (opsin 2), also known as rhodopsin, is expressed in retinal rod photoreceptors and is responsible for vision at low light intensity. Certain variants lead to autosomal recessive or autosomal dominant retinitis pigmentosa and congenital stationary night blindness. OPN2 is a representative animal opsin, one of the first studied. In fact, bovine OPN2 is the first opsin to be sequenced, as well as the first GPCR whose crystal structure was resolved experimentally. Many studies on the functional mechanisms of animal opsins also focus on OPN2. Consequently, I chose OPN2 to conduct a functional analysis in this study in order to further explore the effectiveness of the QTY code in redesigning retinylidene proteins.
OPN3 (opsin 3), also known as encephalopsin or panopsin, is activated by blue and ultraviolet A light. It was discovered in the brain. It is also expressed in melanocytes and keratinocytes in the skin and regulates functions such as melanogenesis, cell differentiation, and glucose uptake. Its expression is also found in the liver, pancreas, kidney, lung, heart, and skeletal muscles. When expressed in neurons, the release of neurotransmitters is inhibited by light, making OPN3 a useful inhibitory optogenetic tool.
OPN4 (opsin 4), also known as melanopsin, is expressed in ipRGC (intrinsically photosensitive retinal ganglion cells) in the ganglion cell layer in the retina. It is essential for non-image-forming responses to light, including the pupillary reflex, optokinetic visual tracking response, and photoentrainment and regulation of circadian rhythm. Certain variants of OPN4 lead to seasonal affective disorder and other circadian rhythm disorders. Rendering OPN4-containing ipRGCs capable of image formation is also a potential pathway for the treatment of various eye diseases such as retinitis pigmentosa and diabetic retinopathy.
OPN5 (opsin 5), also known as neuropsin, is activated by blue and ultraviolet A light. It is expressed in the retina and contributes to the regulation of light-dependent vascular development and photoentrainment in the cornea and retina. Certain variants of OPN5 may lead to cycloplegia, paralysis of the ciliary muscle in the eye.
RGR (RPE-retinal GPCR) is expressed in RPE (retinal pigmented epithelium) and Müller cells in the retina. Unlike the aforementioned human opsins, RGR preferentially binds all-trans-retinal and may catalyze its isomerization into 11-cis-retinal via a retinochrome-like mechanism. It is expressed only in tissue surrounding photoreceptors and plays a role in the light-dependent synthesis of visual chromophore.
RRH (RPE-derived rhodopsin homolog), also known as peropsin, is localized in the microvilli of RPE cells that surround photoreceptor outer segments. It is another protein that preferentially binds to all-trans-retinal. Although not much information is known about RRH, it can reasonably be inferred that it photoisomerizes all-trans-retinal to 11-cis-retinal and may play a role in the upkeep of photoreceptor functions.
Microbial opsins are transmembrane ion pumps or channels. They are also 7TM, though this is due to convergent evolution rather than homology. The chromophore, retinal, usually isomerizes from all-trans to 13-cis, different from the case of animal opsins. In addition, microbial opsins often form oligomers to carry out their functions.
In this study, I select three microbial opsins: BACR (UniProt ID: P02945), BACH (UniProt ID: B0R2U4), and ChR2 (UniProt ID: Q8RUT8).
BACR (bacteriorhodopsin) is a light-driven proton pump. It is one of the first microbial opsins discovered. BACH (halorhodopsin) is a light-driven chloride pump activated by yellow light. ChR2 (channelrhodopsin 2) is a light-activated sodium channel activated by blue light. BACH and ChR2 are among the first optogenetic tools. BACH is used for inhibition, while ChR2 is used for excitation.
This figure shows that, although the QTY code alters the amino acid sequences of the proteins, many of the the protein properties other than water solubility remain the same:
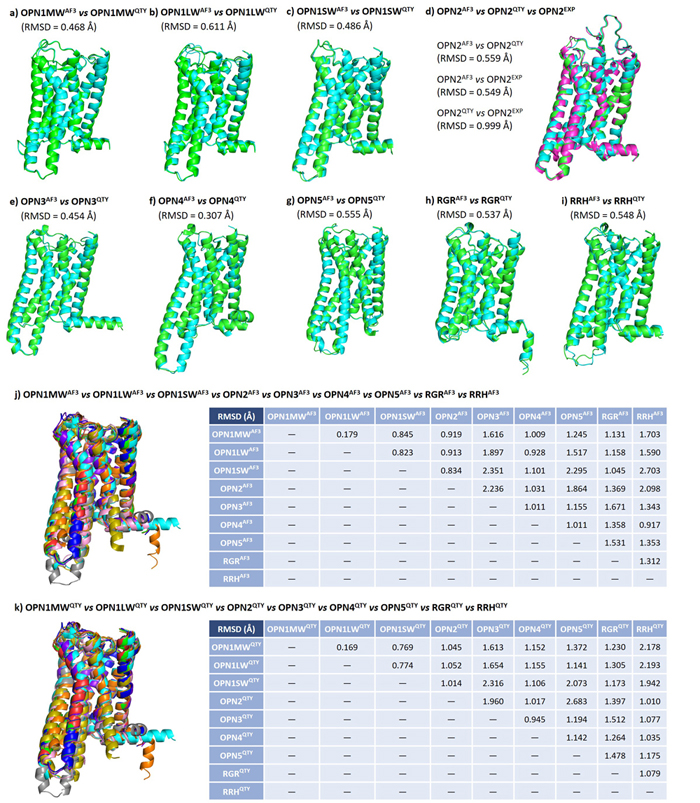
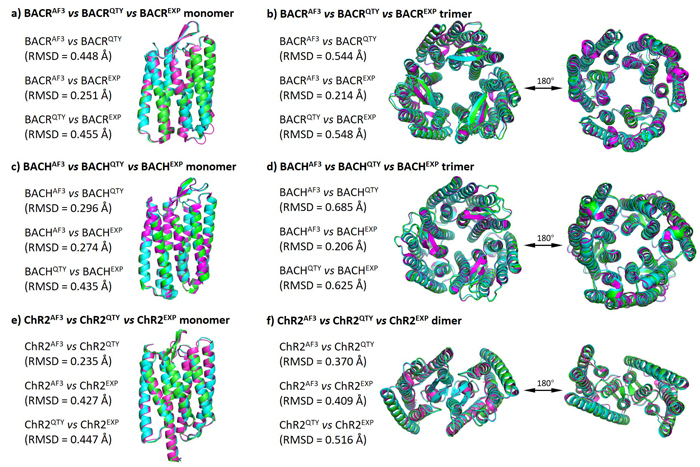
In this figure, I compare the computer-predicted QTY analog structures, computer-predicted native structures, and experimentally-determined native structures of human opsins. It turns out the these three types of structures are very similar to each other. In addition, since all 9 human opsins are evolutionarily related, I perform a pairwise comparison among native and QTY protein structures and found the patterns similar.
In this figure, I compare the computer-predicted QTY analog structures, computer-predicted native structures, and experimentally-determined native structures of microbial opsins. These three types of structures are very similar to each other. Additionally, since these three microbial opsins usually form oligomers (dimers and trimers), I perform a similar comparison between the computer-predicted QTY analog oligomer structures, computer-predicted native oligomer structures, and experimentally-determined native oligomer structures.
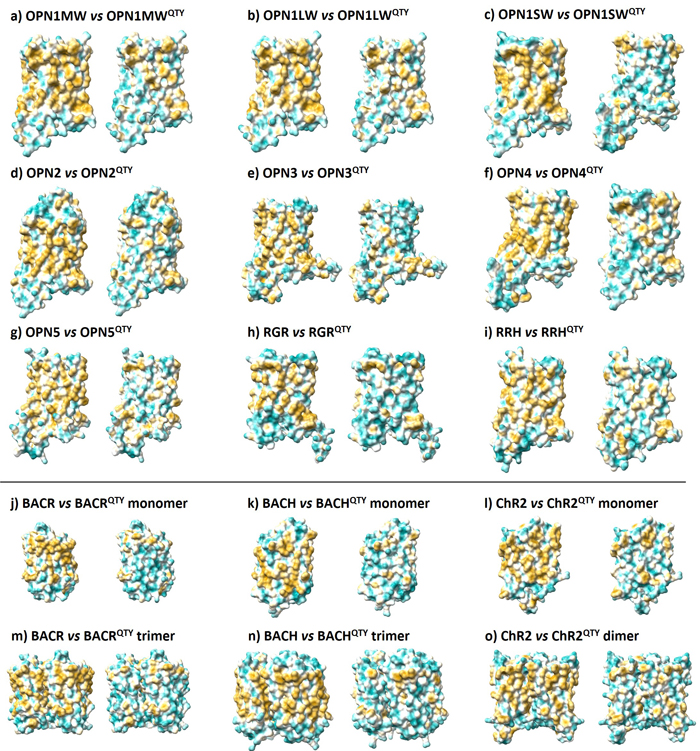
Here, I examine the surface hydrophobicity (degree of water-fearing) in native and QTY structures. Yellow means hydrophobic (water-fearing) and cyan means hydrophilic (water-loving). As observed here, the QTY designed structures have more cyan patches on the surface, so they have higher affinity to water.
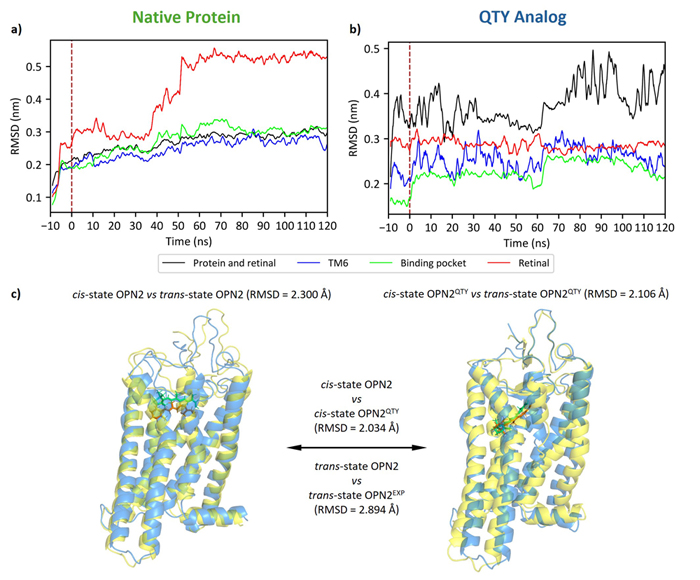
Here, I simulate the 11-cis to all-trans isomerization of the retinal molecule inside rhodopsin (OPN2). This isomerization occurs when a photon hits the protein, i.e., when the protein receives light. As we can deduce from this figure, both the native protein and the QTY analog are likely responsive to light.
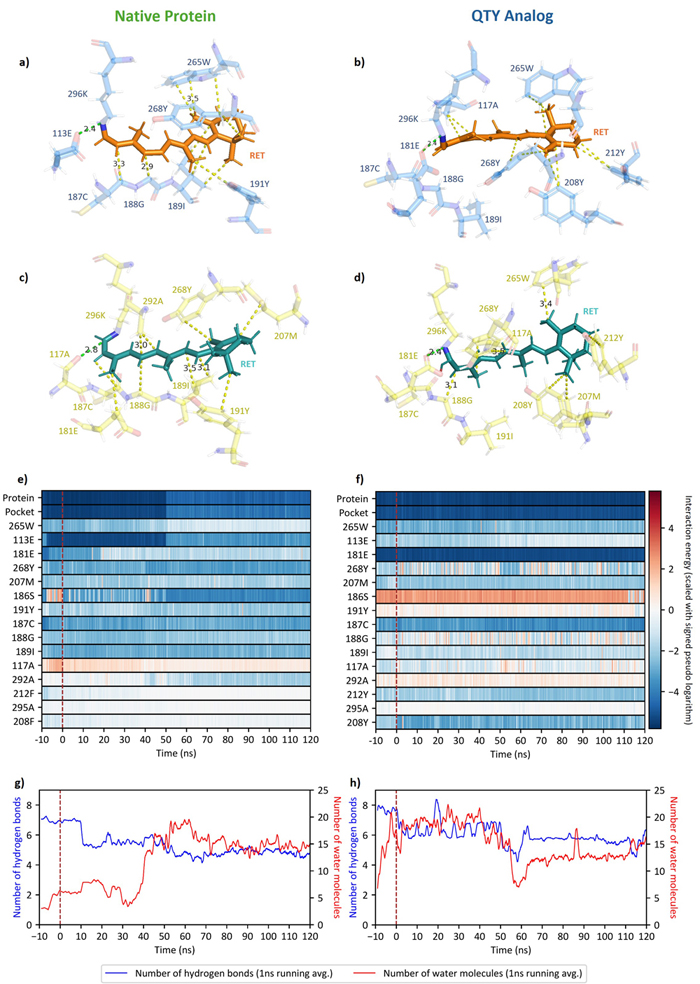
This is a close-up look at the protein's internal machinery and how it responds to light.
During the research, especially when conducting molecular dynamics simulations, I have met several major difficulties:
- At first, I struggled to find reliable parameters for simulating the retinal molecule. I used CGenFF to generate parameters, but the results were not ideal.
- Later, I discovered a file with verified parameters on the NAMD wiki. However, I was using the CHARMM36 force field, where the units and atom naming differed from the NAMD file. This required me to "translate" between the two file formats. Here are some notes I made during the translation:
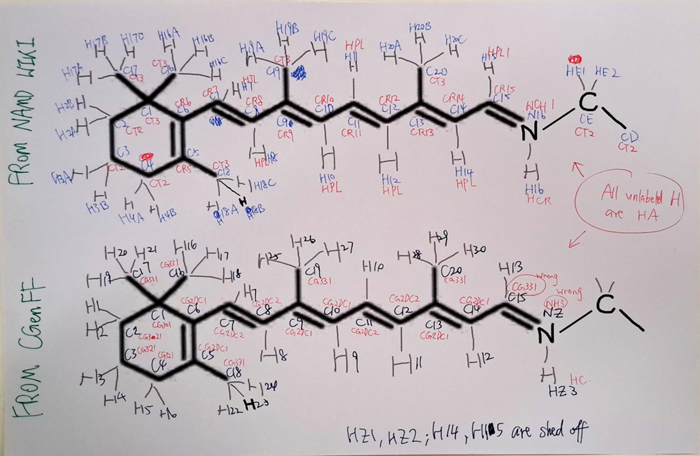
- Simulating the isomerization of retinal was particularly tricky. I had to rotate the molecule manually, but first I needed to determine the correct part to rotate. After consulting multiple papers, I finally found the proper method.
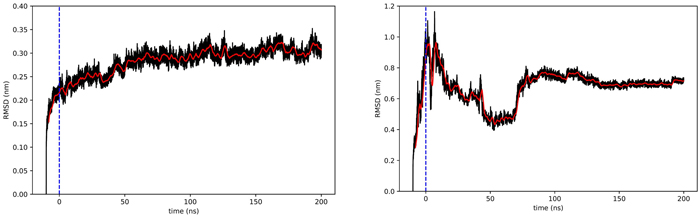
- During simulation, I observed a strange phenomenon. The figure below shows the RMSD (a metric of structural change) for native rhodopsin (left) and the QTY-designed rhodopsin (right). There was a sudden, abrupt change in the QTY version. At first, I was puzzled because it was as if the protein had been struck by an invisible force. I later realized I had omitted an auxiliary “tail” of the protein during visualization. It was exactly this tail that whipped the rest of the structure into a strange shape. Once I removed the tail, the problem was resolved.
- After retinal isomerization, water occasionally entered the protein erroneously, which also took me some time to figure out and resolve.
I also imagine potential future studies and applications:
The findings of this study indicate that the QTY code could be used as a robust tool to design water-soluble retinylidene proteins.
This could, in the first place, facilitate the study of these proteins. The water solubility of QTY analogues makes it easier to purify and investigate proteins, especially those such as RGR and RRH, which have not been studied much but may have considerable clinical relevance. The water solubility also reduces the difficulty in recording the different phases of the photocycle, in comparison with native membrane opsins. Another application may be the rapid design of new optogenetic tools.
Furthermore, water-soluble QTY opsins could be useful clinically, since they could be delivered and/or expressed in the eye without forming aggregates and precipitating. It is not unreasonable to hypothesize that QTY opsins can still pass through the photocycle and interact with downstream signaling proteins. In light of this, QTY opsins may facilitate the optogenetic therapy of ophthalmological diseases, providing a potential pathway in restoring basic vision for those who have lost it.
Finally, water-soluble opsins have the potential to be harvested in mass and may be used to design new biomimetic light-sensing systems, which may have applications in bioengineering.