Development of the MAAM
In 2024, I developed the Mltimodal Abnormity-Aware Model (MAAM) for automatic diagnosis of ophthalmological diseases. In the process, I gained much knowledge in both ophthalmology and deep learning. In this blog, I will introduce the MAAM and key results, share my story of the MAAM and what I learned along the way, discuss difficulties and limitations, and present some notes on the resources I referenced.
Introduction to the MAAM
The Multimodal Abnormity-Aware Model (MAAM) provides reference for both doctors and patients, assisting the diagnosis of ocular diseases. It takes the input of two modalities, i.e., optical coherence tomography (OCT) and color fundus images, and identifies the abnormities present on the images. This is achieved through Abnormity Models, which consist of convolutional neural networks (CNN) and are able to identify 8 OCT abnormities and 11 fundus ones. Then, the model deduces the desease from the predicted abnormities, simulating the decision-making process of ophthalmologists. This is realized through the Diagnosis Model, which consists of multiple networks of single fully-connected layers and is capable of diagnosing 12 diseases. This is the overall structure of the MAAM:
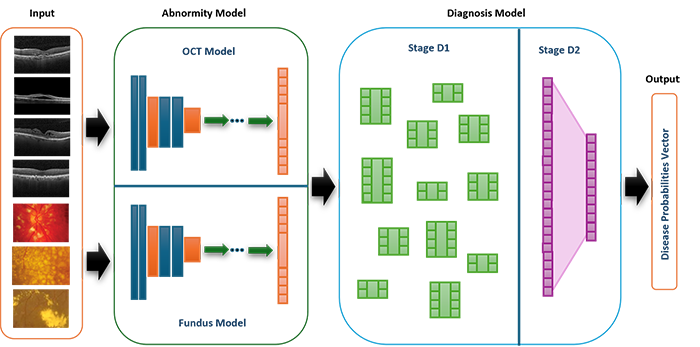
A major advantage of the MAAM is that it partially opens the "black box" of neural networks and makes the results more interpretable through the following methods:
- I employed a fusion mechanism to fuse information from multiple images and different modalities.
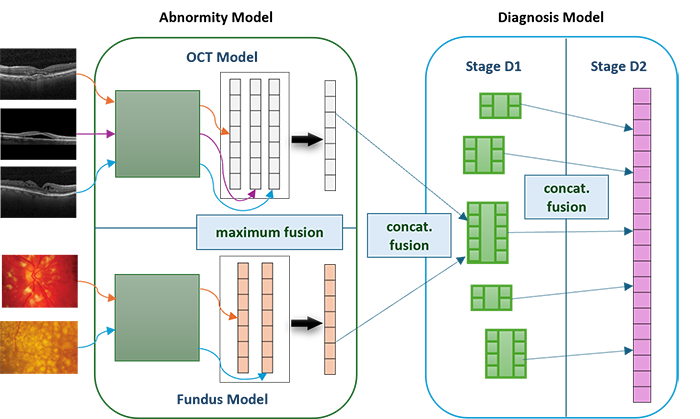
- I leveraged a deduction criteria from abnormities to diseases, simulating the diagnosis process of doctors.
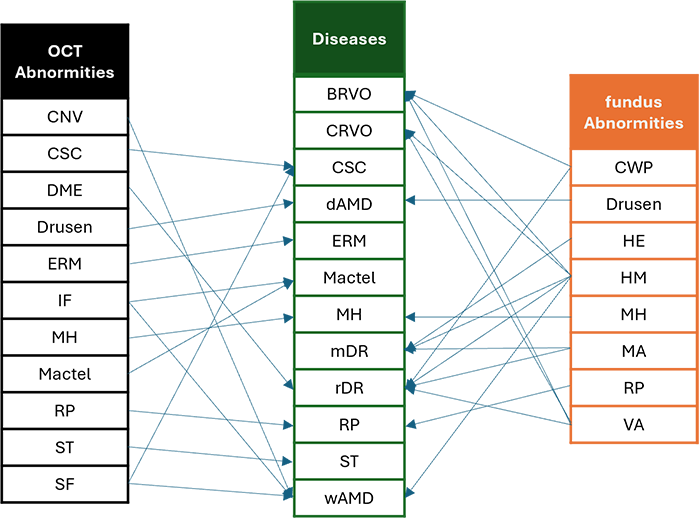
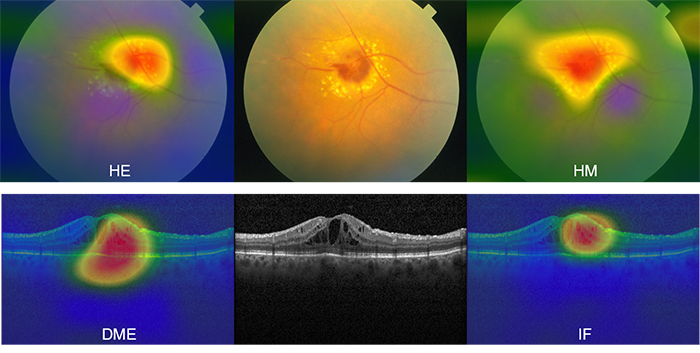
- I added a highlighting function that enables the model to highlight the location of the abnormities on the image.
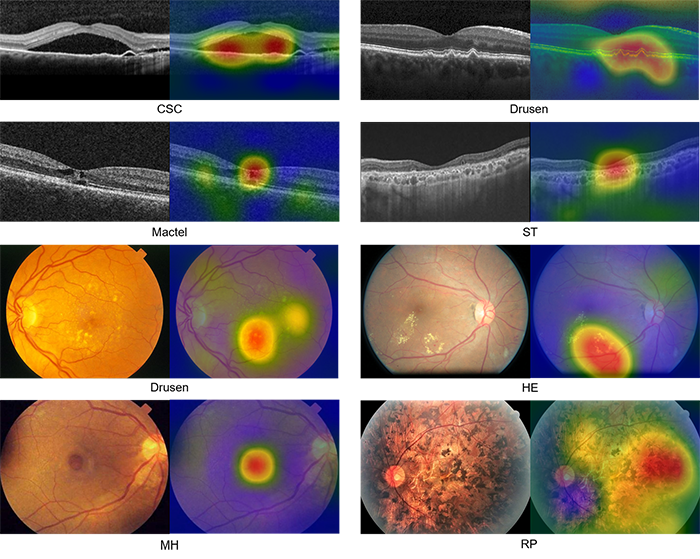
- I introduced codominant abnormities and diseases, allowing the model to suggest multiple possibilities to the user.
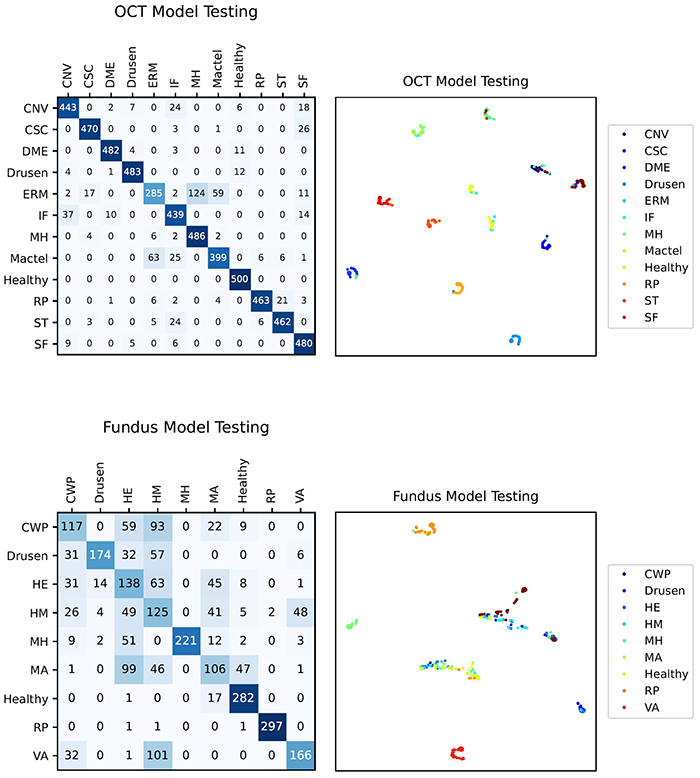
The following figure shows some results (confusion matrices and t-SNE graphs) of the Abnormity models. The performance are not particularly satisfying, especially considering the relatively low accuracy of the Fundus Model.
These are some of the results (accuracy and confusion matrix) of the Diagnosis Model. The Stage D2 of the Diagnosis Model has increased accuracy compared to the Stage D1, which in turn is more accurate than the Abnormity Models.
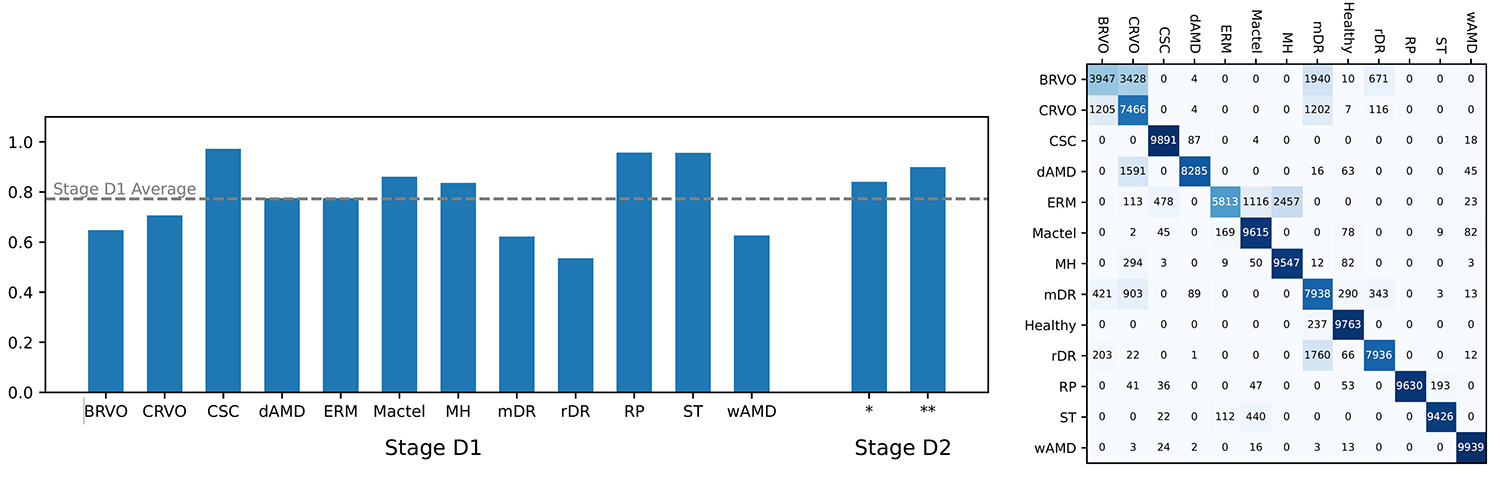
Despite that the data quality is not highly satisfactory, the model still reaches an overall accuracy of 89%, which verifies that the model is likely feasible.
In the future, the MAAM could be re-trained with real clinical data to reach higher accuracy. It is also extensible, allowing the addition of new abnormities and diseases to further enhance its diagnostic capability.
The Story of the MAAM
In 2023, Grandma had an eye examination. I saw many terms, numbers, and graphs on her reports, and I wanted to understand them.
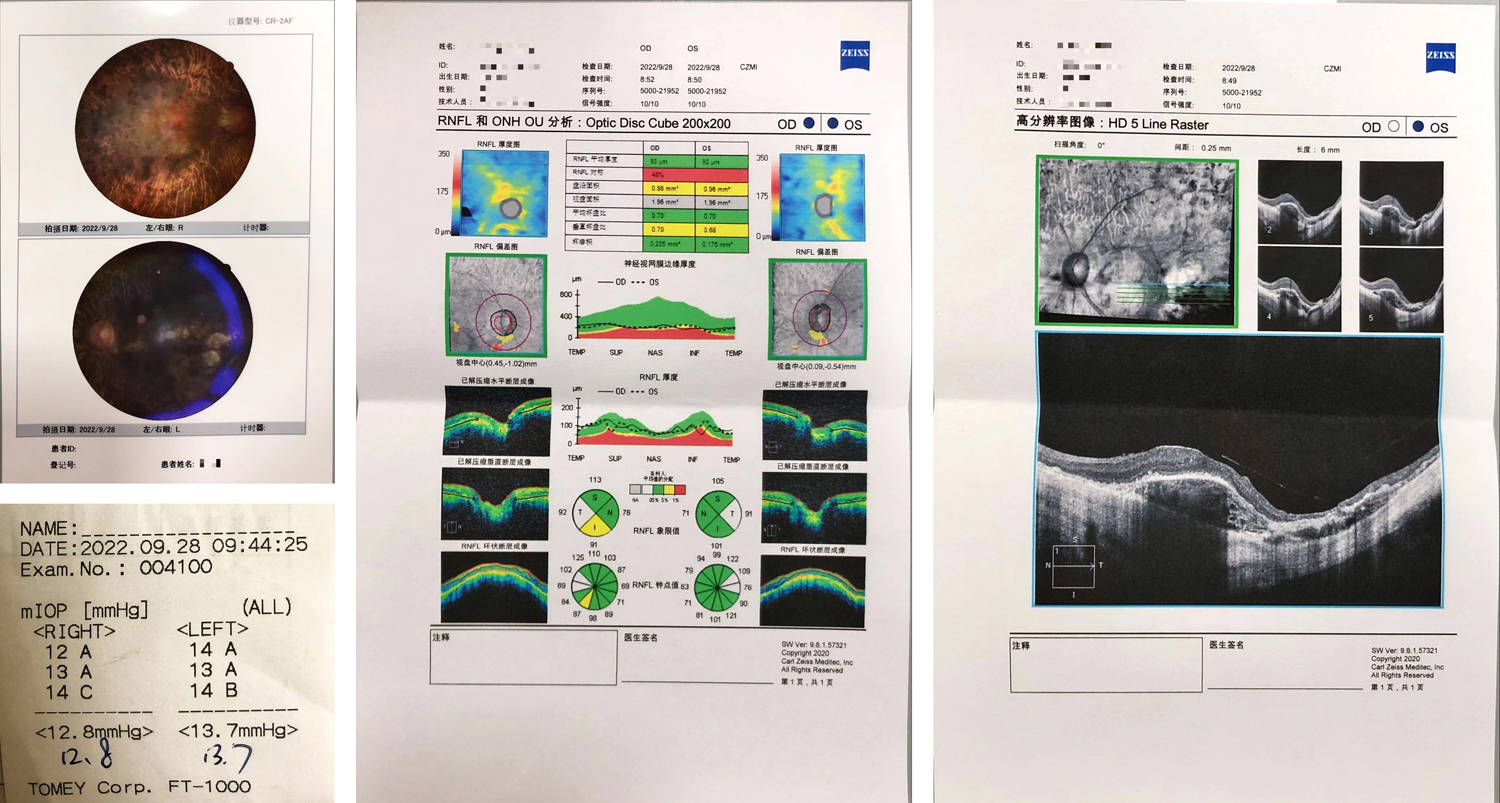
After searching online, I learned about the important terms in the report:
- OD means right eye (oculus dexter), OS means left eye (oculus sinister), and OU means both eyes (oculus uterque).
- NAS means nasal (medial, towards the nose), TEMP means temporal (lateral, towards the temple), SUP means superior (upper, towards the forehead), and INF means inferior (lower, towards the mouth).
- RNFL stands for retinal nerve fiber layer and GCC stands for ganglion cell complex.
- Optic disc, also called optic nerve head (ONH), is the portion of the optic nerve visible in fundus images. Optic cup is the central area of the optic disc that is slightly depressed and lighter in color.
- mIOP stands for mean intraocular pressure, i.e., the average pressure of the fluid inside the eye.
After understanding the terms, I was eager to know what those medical images meant. Therefore, at the start of 2024, I began reading books and teaching myself the principles of ophthalmological diagnosis. I learned about ocular structures, pathological symptoms, as well as the interpretation of optical coherence tomography (OCT) and color fundus images. Some of the resources are recorded in the last section of the blog.
Having read the books, I realized that ophthalmological diagnosis is very complex process. It is difficult for every patient or family to grasp. Moreover, even ophthalmologists may commit mistakes, such as overlooking minute abnormities. Therefore, I considered developing a computer system that could assist the ophthalmological diagnosis and make the results interpretable for patients and families. From February to April, I read papers on ophthalmological diagnosis systems that utilize deep learning. I also learned to implement deep learning models and algorithms with Python (PyTorch), including convolutional neural networks (such as VGG, ResNet, and U-Net), Grad-CAM, and Cycle-GAN.
In April, I started designing my own model. But first, I needed to collect data.
I mainly collected data from online databases. After downloading the images, I made some further modifications. For example, some images contained text annotations. I either edited out the annotation or excluded the image from the dataset. In addition, some images were labeled with only abnormities or only diseases. I then considered the deductive relationship between abnormities and diseases, and labeled these images myself in order to fit the classification scheme required by the MAAM.
After these steps, I still faced the problem that certain rare diseases had only a few images, which caused an imbalance of data. I used search engines to gather standalone images from academic papers and websites. Then, I learned from a paper and used Cycle-GAN, a deep learning method, to generate new images based on the few existing ones. Since some generated images were inaccurate or unusable, I manually filtered all 18,000 images to obtain the final dataset for training MAAM.
From May to August, I wrote the program for the MAAM system, trained it on a desktop computer at home, and wrote a paper summarizing my results, some of which have been introduced above.
Difficulties and Limitations
I encountered some difficulties in the process of gathering data and training models:
- The insufficiency of data was a large issue. I was interested in studying some of the rare diseases, including retinitis pigmentosa, but they had insufficient images. Thankfully, I managed to find the method of Cycle-GAN to generate images and augment the data.
- During the training, the model could not at first reach high accuracy. I tried different convolutional neural networks (ResNet18, ResNet50, ResNet152, and VGG16) and found out the best network for the OCT Model and the Fundus Model, which improved accuracy. I later also found that the Diagnosis Model also boosted accuracy.
- The Abnormity Models often gave wrong predictions at first, but it turned out that the predicted abnormity was actually correct – just that it was not the same with the label, as there was only one label per image. To cope with this, I devised the concept of co-dominant abnormities, which improved accuracy. Later, I expanded this to the Diagnosis Model, introducing co-dominant diseases as well.
In the peer review process, reviewers identified some limitations that I was unable to resolve:
- Most importantly, the data are collected from heterogeneous sources and do not offer rigorous enough conclusions.
- Due to a lack of OCT and Fundus data from the same patient, I was unable to establish direct individual-specific correlations between the two modalities, which may significantly decrease the model's ability to integrate information from the two modalities.
- There are no ophthalmological experts involved in the labelling process, decreasing the accuracy.
- I did not perform a quantitative assessment of how clinically realistic the Cycle-GAN-generated images are. Moreover, the Cycle-GAN-generated images still tend to be similar to the original images, which decreases the variety of the data for rare diseases and may cause over-fitting.
- I only employed relatively simple fusion methods and did not compare them with alternative techniques such as attention-based fusion or gated fusion.
Due to these reasons, especially the first one, my manuscript was not accepted. Nevertheless, it has been a valuable learning opportunity for me. In the future, I would like to study more deep learning techniques to understand more fusion mechanisms and other algorithms.
My Notes on Books, Papers, and Websites
This section contains a list of some of the books, papers, and websites that I have consulted, together with my comments. These fall into three categories: resources with which I learned ophthalmology, papers on automatic diagnosis, and my data sources.
On Ophthalmology
Here are some books and websites on ophthalmology that I studied.
- Agarwal, A., & Jacob, S. (2010). Color atlas of ophthalmology: The quick-reference manual for diagnosis and treatment (2nd ed). Thieme.
- Duker, J. S., Waheed, N. K., & Goldman, D. R. (Eds.). (2022). Handbook of retinal OCT (Second edition). Elsevier.
- Ichhpujani, P., & Thakur, S. (Eds.). (2021). Artificial Intelligence and Ophthalmology: Perks, Perils and Pitfalls. Springer Singapore. https://doi.org/10.1007/978-981-16-0634-2
- Mittal, S. K., & Agarwal, R. K. (2021). Textbook of ophthalmology. Thieme.
- Normal OCT Anatomy. (n.d.). https://en.octclub.org/normal-oct-anatomisi/
- Trucco Emanuele, Tom MacGillivray, & Yanwu Xu. (2019). Computational Retinal Image Analysis Tools, Applications and Perspectives. In Computational Retinal Image Analysis Tools, Applications and Perspectives (pp. i–iii). Elsevier. https://doi.org/10.1016/B978-0-08-102816-2.09989-5
- Wolf, S., Kirchhof, B., & Reim, M. (2006). The ocular fundus: From findings to diagnosis. Thieme.
- This book contains images of diverse ophthalmological conditions, ranging from physical trauma to neurological diseases, including both external photos and specialized medical images. These are accompanied by concise explanations. I was able to learn about the actual "appearance" of various conditions.
- This handbook focuses on OCT, a technique that captures images of the retina, especially cross-sections. I learned about the mechanisms and applications of different types of OCT, as well as how to interpret the most common type, spectral domain OCT. The clinical examples are also great for helping me connect image patterns with specific retinal conditions.
- This book is a collection of reviews on how artificial intelligence has contributed and may contribute to the field of ophthalmology. I learned how algorithms can assist in diagnosing diseases, predicting progression, classifying subtypes, and even guiding treatment decisions. Meanwhile, the book also discussed limitations and ethical considerations, which reminded me that such technology should complement clinical judgement, not replace it – at least for now.
- This textbook provided me with a systematic framework for ophthalmology. Whenever I met a new concept, I referred to the corresponding chapter in this textbook. After reading some sections, I learned the foundational anatomy and physiology of the eye, as well as how certain diseases are classified and managed.
- This is a succinct and beginner-friendly online resource, a key reference for me during early stages of learning. I got to know about the normal retinal structures in OCT images, including retinal layers and key landmarks. Understanding the normal anatomy was essential for my later attempts to interpret abnormal findings, and it gave me confidence when approaching real clinical data.
- Through this book, I learned how we could analyze retinal images through computational methods. I saw techniques for landmark detection, segmentation, and lesion identification in fundus and OCT images, as well as real-world challenges that must be addressed to make automated analysis reliable in clinical practice.
- This book offered me a window into fundus photography, a modality different from OCT. I learned how to carefully examine the fundus and interpret subtle signs that are less noticeable than signs in OCT images.
On Automatic Ophthalmological Diagnosis
These papers are reviews that provided an overview on certain aspects of automatic ophthalmological diagnosis.
- Artificial Intelligence, Machine Learning and Deep Learning in Ophthalmology. (n.d.).
- Daich Varela, M., Sen, S., De Guimaraes, T. A. C., Kabiri, N., Pontikos, N., Balaskas, K., & Michaelides, M. (2023). Artificial intelligence in retinal disease: Clinical application, challenges, and future directions. Graefe's Archive for Clinical and Experimental Ophthalmology, 261(11), 3283–3297. https://doi.org/10.1007/s00417-023-06052-x
- Fumero, F., Alayon, S., Sanchez, J. L., Sigut, J., & Gonzalez-Hernandez, M. (2011). RIM-ONE: An open retinal image database for optic nerve evaluation. 2011 24th International Symposium on Computer-Based Medical Systems (CBMS), 1–6. https://doi.org/10.1109/CBMS.2011.5999143
- Grewal, P. S., Oloumi, F., Rubin, U., & Tennant, M. T. S. (2018). Deep learning in ophthalmology: A review. Canadian Journal of Ophthalmology, 53(4), 309–313. https://doi.org/10.1016/j.jcjo.2018.04.019
- Mitamura, Y., Mitamura-Aizawa, S., Nagasawa, T., Katome, T., Eguchi, H., & Naito, T. (2012). Diagnostic imaging in patients with retinitis pigmentosa. The Journal of Medical Investigation, 59(1,2), 1–11. https://doi.org/10.2152/jmi.59.1
- Moraru, A., Costin, D., Moraru, R., & Branisteanu, D. (2020). Artificial intelligence and deep learning in ophthalmology—Present and future (Review). Experimental and Therapeutic Medicine. https://doi.org/10.3892/etm.2020.9118
- Srivastava, O., Tennant, M., Grewal, P., Rubin, U., & Seamone, M. (2023). Artificial intelligence and machine learning in ophthalmology: A review. Indian Journal of Ophthalmology, 71(1), 11. https://doi.org/10.4103/ijo.IJO_1569_22
- Ting, D. S. W., Pasquale, L. R., Peng, L., Campbell, J. P., Lee, A. Y., Raman, R., Tan, G. S. W., Schmetterer, L., Keane, P. A., & Wong, T. Y. (2019). Artificial intelligence and deep learning in ophthalmology. British Journal of Ophthalmology, 103(2), 167–175. https://doi.org/10.1136/bjophthalmol-2018-313173
- Yanagihara, R. T., Lee, C. S., Ting, D. S. W., & Lee, A. Y. (2020). Methodological Challenges of Deep Learning in Optical Coherence Tomography for Retinal Diseases: A Review. Translational Vision Science & Technology, 9(2), 11. https://doi.org/10.1167/tvst.9.2.11
Besides describing the role of AI in diagnosing different retinal diseases (diabetic retinopathy, age-related macular degeneration, inherited retinal disorders, retinopathy of prematurity, etc.), this paper also provides a brief guide to different types of machine learning and AI models that are commonly used. This provided me with a general idea of what to expect in other papers I read.
The review highlighted that deep learning can support telemedicine, enabling screening, diagnosis, and monitoring in primary care or community settings. To me, this seems particularly important for those who reside in rural areas and are not able to receive timely diagnosis and referral. With good deep learning models, these people would receive better diagnosis and treatment.
These papers pertain to the color fundus modality. They describe models that diagnose diseases from fundus images.
- Burlina, P. M., Joshi, N., Pekala, M., Pacheco, K. D., Freund, D. E., & Bressler, N. M. (2017). Automated Grading of Age-Related Macular Degeneration From Color Fundus Images Using Deep Convolutional Neural Networks. JAMA Ophthalmology, 135(11), 1170. https://doi.org/10.1001/jamaophthalmol.2017.3782
- Choi, J. Y., Yoo, T. K., Seo, J. G., Kwak, J., Um, T. T., & Rim, T. H. (2017). Multi-categorical deep learning neural network to classify retinal images: A pilot study employing small database. PLOS ONE, 12(11), e0187336. https://doi.org/10.1371/journal.pone.0187336
- Decencière, E., Cazuguel, G., Zhang, X., Thibault, G., Klein, J.-C., Meyer, F., Marcotegui, B., Quellec, G., Lamard, M., Danno, R., Elie, D., Massin, P., Viktor, Z., Erginay, A., Laÿ, B., & Chabouis, A. (2013). TeleOphta: Machine learning and image processing methods for teleophthalmology. IRBM, 34(2), 196–203. https://doi.org/10.1016/j.irbm.2013.01.010
- Decencière, E., Zhang, X., Cazuguel, G., Lay, B., Cochener, B., Trone, C., Gain, P., Ordonez, R., Massin, P., Erginay, A., Charton, B., & Klein, J.-C. (2014). FEEDBACK ON A PUBLICLY DISTRIBUTED IMAGE DATABASE: THE MESSIDOR DATABASE. Image Analysis & Stereology, 33(3), 231. https://doi.org/10.5566/ias.1155
- Fang, H., Li, F., Fu, H., Sun, X., Cao, X., Lin, F., Son, J., Kim, S., Quellec, G., Matta, S., Shankaranarayana, S. M., Chen, Y.-T., Wang, C., Shah, N. A., Lee, C.-Y., Hsu, C.-C., Xie, H., Lei, B., Baid, U., … group, iChallenge-A. study. (2022). ADAM Challenge: Detecting Age-related Macular Degeneration from Fundus Images. IEEE Transactions on Medical Imaging, 41(10), 2828–2847. https://doi.org/10.1109/TMI.2022.3172773
- Gargeya, R., & Leng, T. (2017). Automated Identification of Diabetic Retinopathy Using Deep Learning. Ophthalmology, 124(7), 962–969. https://doi.org/10.1016/j.ophtha.2017.02.008
- Gulshan, V., Peng, L., Coram, M., Stumpe, M. C., Wu, D., Narayanaswamy, A., Venugopalan, S., Widner, K., Madams, T., Cuadros, J., Kim, R., Raman, R., Nelson, P. C., Mega, J. L., & Webster, D. R. (2016). Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. JAMA, 316(22), 2402. https://doi.org/10.1001/jama.2016.17216
- Li, B., Chen, H., Zhang, B., Yuan, M., Jin, X., Lei, B., Xu, J., Gu, W., Wong, D. C. S., He, X., Wang, H., Ding, D., Li, X., Chen, Y., & Yu, W. (2021). Development and evaluation of a deep learning model for the detection of multiple fundus diseases based on colour fundus photography. British Journal of Ophthalmology, bjophthalmol-2020-316290. https://doi.org/10.1136/bjophthalmol-2020-316290
- Li, T., Bo, W., Hu, C., Kang, H., Liu, H., Wang, K., & Fu, H. (2021). Applications of Deep Learning in Fundus Images: A Review (No. arXiv:2101.09864). arXiv. http://arxiv.org/abs/2101.09864
- Li, T., Gao, Y., Wang, K., Guo, S., Liu, H., & Kang, H. (2019). Diagnostic assessment of deep learning algorithms for diabetic retinopathy screening. Information Sciences, 501, 511–522. https://doi.org/10.1016/j.ins.2019.06.011
- Masumoto, H., Tabuchi, H., Nakakura, S., Ohsugi, H., Enno, H., Ishitobi, N., Ohsugi, E., & Mitamura, Y. (2019). Accuracy of a deep convolutional neural network in detection of retinitis pigmentosa on ultrawide-field images. PeerJ, 7, e6900. https://doi.org/10.7717/peerj.6900
- Odstrcilik, J., Kolar, R., Budai, A., Hornegger, J., Jan, J., Gazarek, J., Kubena, T., Cernosek, P., Svoboda, O., & Angelopoulou, E. (2013). Retinal vessel segmentation by improved matched filtering: Evaluation on a new high‐resolution fundus image database. IET Image Processing, 7(4), 373–383. https://doi.org/10.1049/iet-ipr.2012.0455
- Orlando, J. I., Fu, H., Breda, J. B., van Keer, K., Bathula, D. R., Diaz-Pinto, A., Fang, R., Heng, P.-A., Kim, J., Lee, J., Lee, J., Li, X., Liu, P., Lu, S., Murugesan, B., Naranjo, V., Phaye, S. S. R., Shankaranarayana, S. M., Sikka, A., … Bogunović, H. (2020). REFUGE Challenge: A Unified Framework for Evaluating Automated Methods for Glaucoma Assessment from Fundus Photographs. Medical Image Analysis, 59, 101570. https://doi.org/10.1016/j.media.2019.101570
- Park, S. J., Shin, J. Y., Kim, S., Son, J., Jung, K.-H., & Park, K. H. (2018). A Novel Fundus Image Reading Tool for Efficient Generation of a Multi-dimensional Categorical Image Database for Machine Learning Algorithm Training. Journal of Korean Medical Science, 33(43), e239. https://doi.org/10.3346/jkms.2018.33.e239
- Son, J., Shin, J. Y., Kong, S. T., Park, J., Kwon, G., Kim, H. D., Park, K. H., Jung, K.-H., & Park, S. J. (2023). An interpretable and interactive deep learning algorithm for a clinically applicable retinal fundus diagnosis system by modelling finding-disease relationship. Scientific Reports, 13(1), 5934. https://doi.org/10.1038/s41598-023-32518-3
- Voets, M., Møllersen, K., & Bongo, L. A. (2019). Reproduction study using public data of: Development and validation of a deep learning algorithm for detection of diabetic retinopathy in retinal fundus photographs. PLOS ONE, 14(6), e0217541. https://doi.org/10.1371/journal.pone.0217541
The ADAM challenge showed the importance of large, well-annotated datasets and standardized evaluation for training and comparing AI models. The authors emphasized the necessity to incorporate professional clinical knowledge and use ensemble strategies to improving deep learning performance.
I learned that deep learning is widely applied to fundus images for tasks like lesion segmentation, biomarker detection, disease diagnosis, and image synthesis. The review highlighted the importance of large, publicly available datasets and careful task-specific evaluation. It also emphasized recognizing common limitations in models and exploring solutions.
In this paper, the authors showed that deep learning can accurately detect multiple retinal abnormities and thus diagnose major eye diseases from fundus images. The study highlighted the importance of explainability, introducing a concept called the counterfactual attribution ratio (CAR) to show how findings of abnormities contribute to predictions of the disease. This idea directly inspired the abnormity-to-disease deduction mechanism in the MAAM.
These papers pertain to the other modality, OCT. They describe models that diagnose diseases from OCT images.
- Camino, A., Wang, Z., Wang, J., Pennesi, M. E., Yang, P., Huang, D., Li, D., & Jia, Y. (2018). Deep learning for the segmentation of preserved photoreceptors on en face optical coherence tomography in two inherited retinal diseases. Biomedical Optics Express, 9(7), 3092. https://doi.org/10.1364/BOE.9.003092
- Chen, T.-C., Lim, W. S., Wang, V. Y., Ko, M.-L., Chiu, S.-I., Huang, Y.-S., Lai, F., Yang, C.-M., Hu, F.-R., Jang, J.-S. R., & Yang, C.-H. (2021). Artificial Intelligence–Assisted Early Detection of Retinitis Pigmentosa—The Most Common Inherited Retinal Degeneration. Journal of Digital Imaging, 34(4), 948–958. https://doi.org/10.1007/s10278-021-00479-6
- Choi, K. J., Choi, J. E., Roh, H. C., Eun, J. S., Kim, J. M., Shin, Y. K., Kang, M. C., Chung, J. K., Lee, C., Lee, D., Kang, S. W., Cho, B. H., & Kim, S. J. (2021). Deep learning models for screening of high myopia using optical coherence tomography. Scientific Reports, 11(1), 21663. https://doi.org/10.1038/s41598-021-00622-x
- Clinically applicable deep learning for diagnosis and referral in retinal disease | Nature Medicine. (n.d.). Retrieved March 9, 2024, from https://www.nature.com/articles/s41591-018-0107-6
- Gholami, P., Roy, P., Parthasarathy, M. K., & Lakshminarayanan, V. (2020). OCTID: Optical coherence tomography image database. Computers & Electrical Engineering, 81, 106532. https://doi.org/10.1016/j.compeleceng.2019.106532
- Kawczynski, M. G., Bengtsson, T., Dai, J., Hopkins, J. J., Gao, S. S., & Willis, J. R. (2020). Development of Deep Learning Models to Predict Best-Corrected Visual Acuity from Optical Coherence Tomography. Translational Vision Science & Technology, 9(2), 51. https://doi.org/10.1167/tvst.9.2.51
- Kermany, D. S., Goldbaum, M., Cai, W., Valentim, C. C. S., Liang, H., Baxter, S. L., McKeown, A., Yang, G., Wu, X., Yan, F., Dong, J., Prasadha, M. K., Pei, J., Ting, M. Y. L., Zhu, J., Li, C., Hewett, S., Dong, J., Ziyar, I., … Zhang, K. (2018). Identifying Medical Diagnoses and Treatable Diseases by Image-Based Deep Learning. Cell, 172(5), 1122-1131.e9. https://doi.org/10.1016/j.cell.2018.02.010
- Khan, Z., Khan, F. G., Khan, A., Rehman, Z. U., Shah, S., Qummar, S., Ali, F., & Pack, S. (2021). Diabetic Retinopathy Detection Using VGG-NIN a Deep Learning Architecture. IEEE Access, 9, 61408–61416. https://doi.org/10.1109/ACCESS.2021.3074422
- Kurmann, T., Yu, S., Márquez-Neila, P., Ebneter, A., Zinkernagel, M., Munk, M. R., Wolf, S., & Sznitman, R. (2019). Expert-level Automated Biomarker Identification in Optical Coherence Tomography Scans. Scientific Reports, 9(1), 13605. https://doi.org/10.1038/s41598-019-49740-7
- Lee, C. S., Baughman, D. M., & Lee, A. Y. (2017). Deep Learning Is Effective for Classifying Normal versus Age-Related Macular Degeneration OCT Images. Ophthalmology Retina, 1(4), 322–327. https://doi.org/10.1016/j.oret.2016.12.009
- Li, F., Chen, H., Liu, Z., Zhang, X., Jiang, M., Wu, Z., & Zhou, K. (2019). Deep learning-based automated detection of retinal diseases using optical coherence tomography images. Biomedical Optics Express, 10(12), 6204. https://doi.org/10.1364/BOE.10.006204
- Li, F., Chen, H., Liu, Z., Zhang, X., & Wu, Z. (2019). Fully automated detection of retinal disorders by image-based deep learning. Graefe's Archive for Clinical and Experimental Ophthalmology, 257(3), 495–505. https://doi.org/10.1007/s00417-018-04224-8
- Liu, T. Y. A., Ling, C., Hahn, L., Jones, C. K., Boon, C. J., & Singh, M. S. (2023). Prediction of visual impairment in retinitis pigmentosa using deep learning and multimodal fundus images. British Journal of Ophthalmology, 107(10), 1484–1489. https://doi.org/10.1136/bjo-2021-320897
- Lu, W., Tong, Y., Yu, Y., Xing, Y., Chen, C., & Shen, Y. (2018). Deep learning-based automated classification of multi-categorical abnormalities from optical coherence tomography images. Translational Vision Science & Technology, 7(6), 41–41.
- M.S, A., B, V., K, S., Kawatra, S., & Vaishnava, A. (2021). Detection of Choroidalneovascularization (Cnv) in Retina Oct Images Using Vgg16 and Densenet Cnn [Preprint]. In Review. https://doi.org/10.21203/rs.3.rs-360517/v1
- Rajagopalan, N., N., V., Josephraj, A. N., & E., S. (2021). Diagnosis of retinal disorders from Optical Coherence Tomography images using CNN. PLOS ONE, 16(7), e0254180. https://doi.org/10.1371/journal.pone.0254180
- Smith, T. B., Parker, M., Steinkamp, P. N., Weleber, R. G., Smith, N., Wilson, D. J., VPA Clinical Trial Study Group, & EZ Working Group. (2016). Structure-Function Modeling of Optical Coherence Tomography and Standard Automated Perimetry in the Retina of Patients with Autosomal Dominant Retinitis Pigmentosa. PLOS ONE, 11(2), e0148022. https://doi.org/10.1371/journal.pone.0148022
- Srinivasan, P. P., Kim, L. A., Mettu, P. S., Cousins, S. W., Comer, G. M., Izatt, J. A., & Farsiu, S. (2014). Fully automated detection of diabetic macular edema and dry age-related macular degeneration from optical coherence tomography images. Biomedical Optics Express, 5(10), 3568. https://doi.org/10.1364/BOE.5.003568
- Yoo, T. K., Choi, J. Y., & Kim, H. K. (2021). Feasibility study to improve deep learning in OCT diagnosis of rare retinal diseases with few-shot classification. Medical & Biological Engineering & Computing, 59(2), 401–415. https://doi.org/10.1007/s11517-021-02321-1
This paper demonstrates that deep learning can accurately categorize OCT images for multiple retinal conditions, achieving performance comparable to or exceeding human experts. The study emphasized the value of large, expertly labeled datasets and rigorous validation methods like 10-fold cross-validation. The MAAM adopted a similar mechanism of cross-validation.
I saw how ensemble deep learning models such as multiple ResNet50 networks can achieve high accuracy in classifying OCT images for multiple retinal diseases, even with limited data. The study enhances interpretability through techniques like occlusion testing. It also showed that integrating clinical history with imaging can further enhance performance.
There are few images available for rare retinal conditions, making it difficult to train models accurately. This study shows that few-shot learning (FSL) with GAN-generated images can overcome data scarcity for rare retinal diseases in OCT diagnosis. The synthetic data helped models learn characteristic features, improving accuracy for rare conditions beyond what traditional deep learning approaches could achieve. This strategy was also adopted by the MAAM.
These papers also relate to OCT images, but they describe models that identify abnormities rather than directly diagnosing diseases.
- Abràmoff, M. D., Lou, Y., Erginay, A., Clarida, W., Amelon, R., Folk, J. C., & Niemeijer, M. (2016). Improved Automated Detection of Diabetic Retinopathy on a Publicly Available Dataset Through Integration of Deep Learning. Investigative Opthalmology & Visual Science, 57(13), 5200. https://doi.org/10.1167/iovs.16-19964
- De Fauw, J., Ledsam, J. R., Romera-Paredes, B., Nikolov, S., Tomasev, N., Blackwell, S., Askham, H., Glorot, X., O'Donoghue, B., Visentin, D., Van Den Driessche, G., Lakshminarayanan, B., Meyer, C., Mackinder, F., Bouton, S., Ayoub, K., Chopra, R., King, D., Karthikesalingam, A., … Ronneberger, O. (2018). Clinically applicable deep learning for diagnosis and referral in retinal disease. Nature Medicine, 24(9), 1342–1350. https://doi.org/10.1038/s41591-018-0107-6
- Fang, L., Cunefare, D., Wang, C., Guymer, R. H., Li, S., & Farsiu, S. (2017). Automatic segmentation of nine retinal layer boundaries in OCT images of non-exudative AMD patients using deep learning and graph search. Biomedical Optics Express, 8(5), 2732. https://doi.org/10.1364/BOE.8.002732
- Fang, L., Wang, C., Li, S., Rabbani, H., Chen, X., & Liu, Z. (2019). Attention to Lesion: Lesion-Aware Convolutional Neural Network for Retinal Optical Coherence Tomography Image Classification. IEEE Transactions on Medical Imaging, 38(8), 1959–1970. https://doi.org/10.1109/TMI.2019.2898414
- Leandro, I., Lorenzo, B., Aleksandar, M., Dario, M., Rosa, G., Agostino, A., & Daniele, T. (2023). OCT-based Deep-learning Models for the Identification of Retinal Key Signs. Scientific Reports, 13(1), 14628. https://doi.org/10.1038/s41598-023-41362-4
- Lee, C. S., Tyring, A. J., Deruyter, N. P., Wu, Y., Rokem, A., & Lee, A. Y. (n.d.). Deep-learning based, automated segmentation of macular edema in optical coherence tomography.
- Liefers, B., Taylor, P., Alsaedi, A., Bailey, C., Balaskas, K., Dhingra, N., Egan, C. A., Rodrigues, F. G., Gonzalo, C. G., Heeren, T. F. C., Lotery, A., Müller, P. L., Olvera-Barrios, A., Paul, B., Schwartz, R., Thomas, D. S., Warwick, A. N., Tufail, A., & Sánchez, C. I. (2021). Quantification of Key Retinal Features in Early and Late Age-Related Macular Degeneration Using Deep Learning. American Journal of Ophthalmology, 226, 1–12. https://doi.org/10.1016/j.ajo.2020.12.034
- Lu, W., Tong, Y., Yu, Y., Xing, Y., Chen, C., & Shen, Y. (2018). Deep learning-based automated classification of multi-categorical abnormalities from optical coherence tomography images. Translational Vision Science & Technology, 7(6), 41–41.
- Muhammad, H., Fuchs, T. J., De Cuir, N., De Moraes, C. G., Blumberg, D. M., Liebmann, J. M., Ritch, R., & Hood, D. C. (2017). Hybrid Deep Learning on Single Wide-field Optical Coherence tomography Scans Accurately Classifies Glaucoma Suspects. Journal of Glaucoma, 26(12), 1086. https://doi.org/10.1097/IJG.0000000000000765
- Rasti, R., Rabbani, H., Mehridehnavi, A., & Hajizadeh, F. (2017). Macular OCT classification using a multi-scale convolutional neural network ensemble. IEEE Transactions on Medical Imaging, 37(4), 1024–1034.
- Roy, A. G., Conjeti, S., Karri, S. P. K., Sheet, D., Katouzian, A., Wachinger, C., & Navab, N. (2017). ReLayNet: Retinal layer and fluid segmentation of macular optical coherence tomography using fully convolutional networks. Biomedical Optics Express, 8(8), 3627. https://doi.org/10.1364/BOE.8.003627
- Schlegl, T., Waldstein, S. M., Bogunovic, H., Endstraßer, F., Sadeghipour, A., Philip, A.-M., Podkowinski, D., Gerendas, B. S., Langs, G., & Schmidt-Erfurth, U. (2018). Fully Automated Detection and Quantification of Macular Fluid in OCT Using Deep Learning. Ophthalmology, 125(4), 549–558. https://doi.org/10.1016/j.ophtha.2017.10.031
- Xu, X., Lee, K., Zhang, L., Sonka, M., & Abràmoff, M. D. (2015). Stratified Sampling Voxel Classification for Segmentation of Intraretinal and Subretinal Fluid in Longitudinal Clinical OCT Data. IEEE Transactions on Medical Imaging, 34(7), 1616–1623. IEEE Transactions on Medical Imaging. https://doi.org/10.1109/TMI.2015.2408632
I saw how deep learning can help analyze three-dimensional OCT scans, not just two-dimensional images. This study highlighted the value of tissue segmentation in allowing models to generalize across different imaging devices. Unfortunately, I was not able to obtain first-hand data during the development of the MAAM, and thus was not able to include 3D scans.
I learned that focusing on lesion-specific regions can significantly improve OCT classification. The study introduced a lesion-aware convolutional neural network (LACNN) that uses attention maps to guide the model, mimicking how ophthalmologists focus on local lesions.
This study used binary deep learning models to accurately detect multiple retinal abnormalities from OCT images. It emphasized the value of expert-labeled datasets and showed that efficient labeling strategies can reduce dataset creation time. Indeed, as I found later, labeling strategies were very important for coping with large datasets.
Some models are multimodal, meaning that they take different types of information into consideration – OCT images, fundus images, doctors' descriptions, patient history, etc. Here are some papers that describe multimodal models.
- Andrearczyk, V., & Müller, H. (2018). Deep Multimodal Classification of Image Types in Biomedical Journal Figures. In P. Bellot, C. Trabelsi, J. Mothe, F. Murtagh, J. Y. Nie, L. Soulier, E. SanJuan, L. Cappellato, & N. Ferro (Eds.), Experimental IR Meets Multilinguality, Multimodality, and Interaction (Vol. 11018, pp. 3–14). Springer International Publishing. https://doi.org/10.1007/978-3-319-98932-7_1
- Arcadu, F., Benmansour, F., Maunz, A., Michon, J., Haskova, Z., McClintock, D., Adamis, A. P., Willis, J. R., & Prunotto, M. (2019). Deep Learning Predicts OCT Measures of Diabetic Macular Thickening From Color Fundus Photographs. Investigative Opthalmology & Visual Science, 60(4), 852. https://doi.org/10.1167/iovs.18-25634
- Bidwai, P., Gite, S., Gupta, A., Pahuja, K., & Kotecha, K. (2024). Multimodal dataset using OCTA and fundus images for the study of diabetic retinopathy. Data in Brief, 52, 110033. https://doi.org/10.1016/j.dib.2024.110033
- Chen, Z., Zheng, X., Shen, H., Zeng, Z., Liu, Q., & Li, Z. (2019). Combination of Enhanced Depth Imaging Optical Coherence Tomography and Fundus Images for Glaucoma Screening. Journal of Medical Systems, 43(6), 163. https://doi.org/10.1007/s10916-019-1303-8
- Farsiu, S., Chiu, S. J., O'Connell, R. V., Folgar, F. A., Yuan, E., Izatt, J. A., & Toth, C. A. (2014). Quantitative Classification of Eyes with and without Intermediate Age-related Macular Degeneration Using Optical Coherence Tomography. Ophthalmology, 121(1), 162–172. https://doi.org/10.1016/j.ophtha.2013.07.013
- Hassan, T., Akram, M. U., Hassan, B., Nasim, A., & Bazaz, S. A. (2015). Review of OCT and fundus images for detection of Macular Edema. 2015 IEEE International Conference on Imaging Systems and Techniques (IST), 1–4. https://doi.org/10.1109/IST.2015.7294517
- Jin, K., Yan, Y., Chen, M., Wang, J., Pan, X., Liu, X., Liu, M., Lou, L., Wang, Y., & Ye, J. (2022). Multimodal deep learning with feature level fusion for identification of choroidal neovascularization activity in age‐related macular degeneration. Acta Ophthalmologica, 100(2). https://doi.org/10.1111/aos.14928
- Lim, W. S., Ho, H.-Y., Ho, H.-C., Chen, Y.-W., Lee, C.-K., Chen, P.-J., Lai, F., Jang, J.-S. R., & Ko, M.-L. (2022). Use of multimodal dataset in AI for detecting glaucoma based on fundus photographs assessed with OCT: Focus group study on high prevalence of myopia. BMC Medical Imaging, 22(1), 206. https://doi.org/10.1186/s12880-022-00933-z
- Mahmudi, T., Kafieh, R., Rabbani, H., Mehri Dehnavi, A., & Akhlagi, M. (2014). Comparison of macular OCTs in right and left eyes of normal people (R. C. Molthen & J. B. Weaver, Eds.; p. 90381W). https://doi.org/10.1117/12.2044046
- RAG-FW: A hybrid convolutional framework for the automated extraction of retinal lesions and lesion-influenced grading of human retinal pathology. (n.d.).
- Thakoor, K. A., Yao, J., Bordbar, D., Moussa, O., Lin, W., Sajda, P., & Chen, R. W. S. (2022). A multimodal deep learning system to distinguish late stages of AMD and to compare expert vs. AI ocular biomarkers. Scientific Reports, 12(1), 2585. https://doi.org/10.1038/s41598-022-06273-w
- Tseng, V. S., Chen, C.-L., Liang, C.-M., Tai, M.-C., Liu, J.-T., Wu, P.-Y., Deng, M.-S., Lee, Y.-W., Huang, T.-Y., & Chen, Y.-H. (2020). Leveraging Multimodal Deep Learning Architecture with Retina Lesion Information to Detect Diabetic Retinopathy. Translational Vision Science & Technology, 9(2), 41. https://doi.org/10.1167/tvst.9.2.41
- Vrochidis, S., Huet, B., Chang, E., & Kompatsiaris, I. (2019). Big Data Analytics for Large‐Scale Multimedia Search (1st ed.). Wiley. https://doi.org/10.1002/9781119376996
- Xu, Z., Wang, W., Yang, J., Zhao, J., Ding, D., He, F., Chen, D., Yang, Z., Li, X., Yu, W., & Chen, Y. (2021). Automated diagnoses of age-related macular degeneration and polypoidal choroidal vasculopathy using bi-modal deep convolutional neural networks. British Journal of Ophthalmology, 105(4), 561–566. https://doi.org/10.1136/bjophthalmol-2020-315817
- Yi, S., Zhang, G., Qian, C., Lu, Y., Zhong, H., & He, J. (2022). A Multimodal Classification Architecture for the Severity Diagnosis of Glaucoma Based on Deep Learning. Frontiers in Neuroscience, 16, 939472. https://doi.org/10.3389/fnins.2022.939472
- Yoo, T. K., Choi, J. Y., Seo, J. G., Ramasubramanian, B., Selvaperumal, S., & Kim, D. W. (2019). The possibility of the combination of OCT and fundus images for improving the diagnostic accuracy of deep learning for age-related macular degeneration: A preliminary experiment. Medical & Biological Engineering & Computing, 57(3), 677–687. https://doi.org/10.1007/s11517-018-1915-z
I saw how multimodal deep learning combining OCT and OCTA can achieve expert-level accuracy in detecting choroidal neovascularization in age-related macular degeneration. The study highlighted the effectiveness of feature-level fusion (FLF) to integrate different imaging modalities. I learned from this idea in developing the MAAM.
This study showed that bi-modal deep learning combining fundus and OCT images can accurately classify age-related macular degeneration and polypoidal choroidal vasculopathy, sometimes surpassing human experts. The study highlighted the importance of integrating complementary imaging modalities and validated model performance against expert diagnoses.
This study combines OCT and fundus images in a multimodal deep learning model, significantly improving diagnostic accuracy for age-related macular degeneration compared to using a single modality. This showed that integrating complementary data sources could help achieve more precise and reliable retinal disease detection.
Finally, another group of papers focuses on retinal layers segmentation, i.e., automatically identifying the different retinal layers in OCT images. Oftentimes, the thickness and evenness of these layers are important for making diagnostic decisions. I did not focus on this because it requires more complex labeling as well as more expertise, both of which were beyond my reach.
- Bagci, A. M., Shahidi, M., Ansari, R., Blair, M., Blair, N. P., & Zelkha, R. (2008). Thickness Profiles of Retinal Layers by Optical Coherence Tomography Image Segmentation. American Journal of Ophthalmology, 146(5), 679-687.e1. https://doi.org/10.1016/j.ajo.2008.06.010
- Burke, J., Engelmann, J., Hamid, C., Reid-Schachter, M., Pearson, T., Pugh, D., Dhaun, N., King, S., MacGillivray, T., Bernabeu, M. O., Storkey, A., & MacCormick, I. J. C. (2023). An open-source deep learning algorithm for efficient and fully-automatic analysis of the choroid in optical coherence tomography (No. arXiv:2307.00904). arXiv. https://doi.org/10.48550/arXiv.2307.00904
- Cheong, K. X., Lim, L. W., Li, K. Z., & Tan, C. S. (2018). A novel and faster method of manual grading to measure choroidal thickness using optical coherence tomography. Eye, 32(2), 433–438. https://doi.org/10.1038/eye.2017.210
- Chiu, S. J., Li, X. T., Nicholas, P., Toth, C. A., Izatt, J. A., & Farsiu, S. (2010). Automatic segmentation of seven retinal layers in SDOCT images congruent with expert manual segmentation. Optics Express, 18(18), 19413. https://doi.org/10.1364/OE.18.019413
- Kromer, R. (2017). An Approach for Automated Segmentation of Retinal Layers In Peripapillary Spectralis SD-OCT Images Using Curve Regularisation. ARCHIVOS DE MEDICINA, 1(2).
- Optical coherence tomography as film thickness measurement technique. (n.d.).
- Payne, J. F., Bruce, B. B., & Yeh, S. (2012). Author Response: Retinal Thickness Measurement in OCT. Investigative Opthalmology & Visual Science, 53(2), 854. https://doi.org/10.1167/iovs.12-9527
Data Sources
I collected data from these databases, including OCT and fundus images together with their labels.
- 1000 Fundus images with 39 categories. (n.d.). Retrieved August 9, 2024, from https://www.kaggle.com/datasets/linchundan/fundusimage1000
- Githinji, P. B., Zhao, K., Wang, J., & Qin, P. (2024). IRFundusSet: An Integrated Retinal Fundus Dataset with a Harmonized Healthy Label (No. arXiv:2402.11488). arXiv. http://arxiv.org/abs/2402.11488
- Kermany, D., Zhang, K., & Goldbaum, M. (2018). Large Dataset of Labeled Optical Coherence Tomography (OCT) and Chest X-Ray Images. 3. https://doi.org/10.17632/rscbjbr9sj.3
- MAFFRE, G. P., Gervais GAUTHIER, Bruno LAY, Julien ROGER, Damien ELIE, Mélanie FOLTETE, Arthur DONJON, Hugo. (n.d.-a). E-ophtha. ADCIS. Retrieved March 23, 2024, from https://www.adcis.net/en/third-party/e-ophtha/
- MAFFRE, G. P., Gervais GAUTHIER, Bruno LAY, Julien ROGER, Damien ELIE, Mélanie FOLTETE, Arthur DONJON, Hugo. (n.d.-b). Messidor. ADCIS. Retrieved March 23, 2024, from https://www.adcis.net/en/third-party/messidor/
- Odstrcilik, J., Kolar, R., Budai, A., Hornegger, J., Jan, J., Gazarek, J., Kubena, T., Cernosek, P., Svoboda, O., & Angelopoulou, E. (2013). Retinal vessel segmentation by improved matched filtering: Evaluation on a new high‐resolution fundus image database. IET Image Processing, 7(4), 373–383. https://doi.org/10.1049/iet-ipr.2012.0455
- Porwal, P. (2018). Indian Diabetic Retinopathy Image Dataset (IDRiD) [Dataset]. IEEE. https://ieee-dataport.org/open-access/indian-diabetic-retinopathy-image-dataset-idrid
- The STARE Project. (n.d.). Retrieved March 23, 2024, from https://cecas.clemson.edu/~ahoover/stare/
- Yoo, T. (2020). Data for: Improved accuracy in OCT diagnosis of rare retinal disease using few-shot learning with generative adversarial networks. 2. https://doi.org/10.17632/btv6yrdbmv.2
